Boolean Operators (AND, OR, NOT) are tools for combining search terms and are inherent part of online database searching. While experienced searchers will use Boolean Operators directly in their search strategies, even novice searchers that just enter a string of terms into a database’s search box will end up indirectly using the Boolean operator AND, as each space between words will be treated by the database as AND, thus combining each term together into a search strategy that would retrieve results that have all terms present.
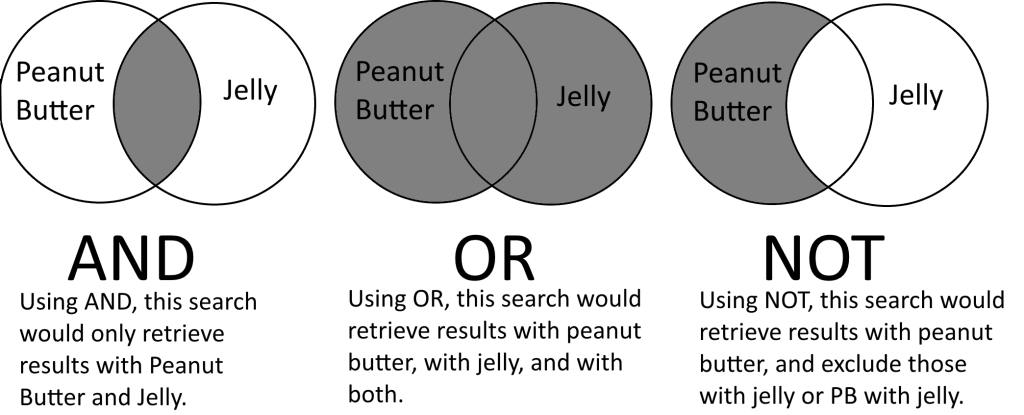
Most search strategies will either use just AND or a combination of both AND and OR. The third Boolean operator, NOT, is much more complicated and requires some understanding to use properly in a search.
Using the Boolean Operator NOT
The Boolean operator NOT can be used when a term or terms needs to be excluded from your search strategy.
For example, if you were interested in articles that looked at children with cancer, but you did not want articles that looked specifically at infants, you could create a search strategy like this:
cancer AND child* NOT infant*
— or —
(cancer AND child*) NOT infant*
The Problem with NOT
When using the Boolean operator NOT to exclude terms, it can become problematic when the database excludes records that contain both the term(s) you want to exclude and the term(s) you want in your search.
In the above example, not only articles about cancer in infants will be excluded from the results but it will also exclude any articles about cancer in both children and infants.
Information professionals (librarians and informationists) advise using the Boolean operator NOT with extreme caution when conducting searches. It’s better to reach out to an information professional for assistance with complex search techniques and how to best proceed with a search when there is a term you want to avoid.
Variations Across Databases
Not all databases function the same way, and using the Boolean operator NOT is no different. While most databases allow for using simply NOT to exclude terms, depending on the database or platform, you might need to use the operator AND NOT instead (Scopus), or once the search is performed use the Exclude button found within the Refine Search panel (also in Scopus).
Takeaway
The Boolean operator NOT should be used with extreme caution. It is best to consult a Librarian on its use in your search.